- Sponsor
- Department of Biochemistry (HOST: Dr. Auinash Kalsotra)
- Speaker
- Dr. Jeongsik Yong
- Contact
- Cara Day
- caraday@illinois.edu
- Phone
- 217-333-2013
- Views
- 212
- Originating Calendar
- Biochemistry Department Seminars
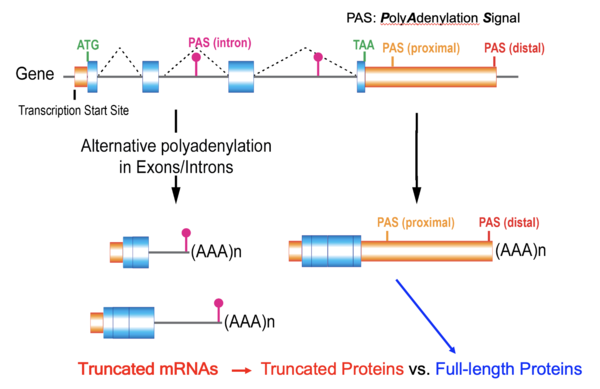
Alternative cleavage and polyadenylation (APA) in introns causes premature transcriptional termination and produces shorter mRNA isoforms from genes. Thus, intronic APA drives the production of shorter protein isoforms in the proteome, but the physiological relevance of intronic APA is elusive. Using APA-targeted profiling, we found that intronic APA events in the transcriptome show a bisectional pattern: a group of genes increase intronic APA while others decrease intronic APA upon changes in cellular mammalian target of rapamycin activity. This discrete pattern of intronic APA persists in normal tissues and tumor samples across 10 examined cancer types in The Cancer Genome Atlas (TCGA) transcriptome datasets. In silico interrogations on the programming of Pfam domains using highly conserved intronic APA events across cancers led to unexpected findings: intronic APA rearranges the truncation of specific Pfam domains among the same family of proteins depending on disease conditions and thus, selectively switches on and off specific Pfam domains’ function without the help of differential gene expression. Intronic APA also features a disease condition-associated regulation of protein length and renders a multifaceted C-terminome in cancer which could be pathogenic. Studies on JMJD6 intronic APA isoform-specific functions using genome engineering suggested that multifaceted pathogenic C-terminone exists in the cancer proteome. Unlike the prevalent shortening of transcripts by 3’-untranslated region APA in the cancer transcriptome, we show that intronic APA is a discrete molecular signature in both normal tissue and cancer transcriptomes and suggest that the dynamics of intronic APA renders a multifunctional C-terminome in the cancer proteome.